Research Article
VDR Gene Polymorphisms and the Risk of Type 1 Diabetes Mellitus: Plots from Meta-Analysis Studies
4961
Views & Citations3961
Likes & Shares
Type 1 diabetes mellitus (T1DM) is a complex autoimmune disease, caused by autoimmune destruction of the insulin secreting b-cells of the islets of Langerhans, in which the immune system plays a key role. The effect of active vitamin D on the pathogenesis of T1DMis directly via the nuclear vitamin D receptor (VDR), which is involved in triggering the immune system against the islet cells of the pancreas. The VDR big-four-inventory polymorphisms (SNPs), FokI (rs2228570), BsmI (rs1544410), ApaI (rs7975232), and TaqI (rs731236), were extensively investigated among T1DM patients from different populations, with inconsistent outcomes. In this review, studies from meta-analysis were merely considered to figure out the role of these SNPs on the risk of T1DM in- and between- populations, taking in consideration their advantages over the individual studies, in terms of accuracy and less bias. Notwithstanding, the results from different continents and between populations was inconclusive, and warrant further studies encompass large sample sizes, equal geographic distributions, tackling haplotypes structure and LD between polymorphisms, and using advanced techniques other than the RFLP for determining individual polymorphisms. Taking these approaches could likely help to emphasize the link between the reviewed SNPs with different functional alleles that may be found in the VDR gene or in genes nearby VDR, and to construct haplotype maps for the candidate genes.
Keywords: Type 1 diabetes, Vitamin D, VDR Polymorphisms
INTRODUCTION
Type 1 Diabetes Worldwide
Type 1 diabetes mellitus (T1DM) is a complex autoimmune disease characterized by reduction or ineffective production of insulin. T1DM is caused by autoimmune destruction of the insulin secreting b-cells of the islets of Langerhans, and is potentially triggered by the immune system, the environmental factors, and many genetics [1-3].
Worldwide, the annual incidence rate of T1DM is approximately 3%, with a proportion of 1 in every 300 persons, and is steadily rising in frequency [4]. Geographically, the incidence of T1DM varies widely, with European (EUR) region having the highest incidence rate (>25 % of the global total prevalence) [2,5,6], followed by the North America and Caribbean (NAC) region (20%), and Western Pacific (WP) region (9%). China and Africa regions reported the lowest incidence rates (1% and 2%, respectively) [5-7]. In the childhood, T1DM is considered the most common chronic disease that constitutes about 5% to 10% of all diabetes cases, with an incidence rate higher in the age between 0-14 years [2,8], and particularly among children less than 5 years of age [9]. It has been estimated that, around 600,900 children were living with diabetes, and almost 98,200 new cases were under the 15 years’ age [5].
Pathogenesis of T1DM
The pathogenesis of T1DM is precisely unknown, despite the supporting evidence linked the damaged b-cells with the releases of b-cells autoantigens [10]. These autoantigens are proposed to be taken up and presented by antigen presenting cells (APCs) to the naïve CD4+ T-helper (Th0) cells in the pancreatic lymph nodes [11]. The IL-12 secreted by the APCs triggers the differentiation of naïve Th0 to active CD4+ T-helper (Th1) cells(pro-inflammatory) that are sequentially release the pro-inflammatory cytokines, IFN-g and IL-2 [10]. The Th1-released cytokines trigger the classical activation of macrophages (CAMs) to release toxic cytokines (TNF-a and IL-1) and free radicals (NO), and in differentiation of CD8+ cytotoxic T cells, collectively to produce inflammatory substances and mediators to execute theb-cells destruction [12-14]. The destruction of the b-cells can be prevented via the setting up of the Th2 cytokines, IL-4 and IL-10. Since, the secretion of IL-4 by natural killer cells (NKs) stimulate the Th0 to differentiate favorably into Th2cells (anti-inflammatory). Further secretion of IL-4 from Th2 cells, in addition to IL-10, prevent Th1 proliferation and macrophage (MF) activation [11,13,15]. The counterbalance of Th1 and Th2 cytokines are the key factors in controlling the pathogenesis in the pancreatic milieu, as well as, in other organ-specific autoimmunity diseases [16]. The role of vitamin D and its counterpart receptor (VDR) were found to be under the path of these settings.
Vitamin D Metabolism and Modulation in T1DM
Vitamin D is a fat-soluble steroid hormone, mainly taken from exposure to sunlight and adsorption through the skin. The biologically active form of vitamin D, the 1,25-dihydroxyvitamin D3 [1,25(OH)2D3] known as calcitriol, is generated from a cascade of reactions, starting from the formation of pre-vitamin D3andvitamin D3(cholecalciferol) from 7-dehydrocholesterol in the skin. Vitamin D3undergoes hydroxylation in the liver by the enzyme vitamin D 25a-hydroxylase (25α-OHase; encoded by CYP2R1) forming25-hydroxyvitamin D3 [25(OH)D3], which is the biologically inactive form of vitamin D3 known as calcidiol. The inactive calcidiol subsequently converted to the active calcitriol in the kidney by 1-alpha hydroxylase (1α-OHase; encoded by CYP27B1) [17-19].
Calcitriol is crucially carried in plasma by vitamin D binding protein (VDBP), which is further facilitate calcitriol cellular endocytosis [20]. Once calcitriol enters the cell, it binds vitamin D receptor (VDR), and subsequent conformation change take place allows the formation of heterodimerization complex with retinoid X receptor (RXR), 1,25(OH)2D3-RXR-VDR. The 1,25(OH)2D3-RXR-VDR complex then cross the nuclear envelop and enter the nucleus, where it can bind to specific DNA sequence elements such as vitamin D response elements (VDRE) located in vitamin D responsive genes [19].
In T1DM, calcitriol has the ability to affect the immune cells directly through binding to the intracellular vitamin D receptor (VDR), which is expressed in a variety of immune cells, including MF, dendritic cells (DCs), and activated T cells [21]. It has been stated that supplementation of calcitriol can significantly inhibit the maturation of DCs to present surface major histocompatibility complex (MHC) class II that can affect Th1 and Th17 differentiation [22]. Furthermore, calcitriol can reduce the ability of MF to present antigens to T cells by decreasing its surface expression to MHC class II [23,24]. It also inhibits the CD8+ T cells to secrete less IFN-g, IL-17, TNF-a, and IL-21, and more TGF-b and IL-5 [21]. Further, calcitriol promotes CD4+ T cells to produce pro-inflammatory cytokines such as IFN-g, IL-17 and IL-21 [25,26]. Vitamin D deficiency and/or a defect in VDR could unlikely increase the Th1/Th2 ratio. The imbalance in the immune response as a risk factor for developing T1DM is not only depend on vitamin D insufficiency, resulted from the sunlight exposure, dietary behaviors, and supplementation intake, but also on polymorphisms in genes involved in the metabolic pathway of vitamin D, including VDR. Collectively, the immunomodulatory effect of calcitriol against autoimmune diseases, particularly T1DM, depends on its ability to induces immunological tolerance and dampen all mechanisms connected with the adaptive immunity that directed the anti-inflammatory effects.
VDR Polymorphisms and the Risk of T1DM
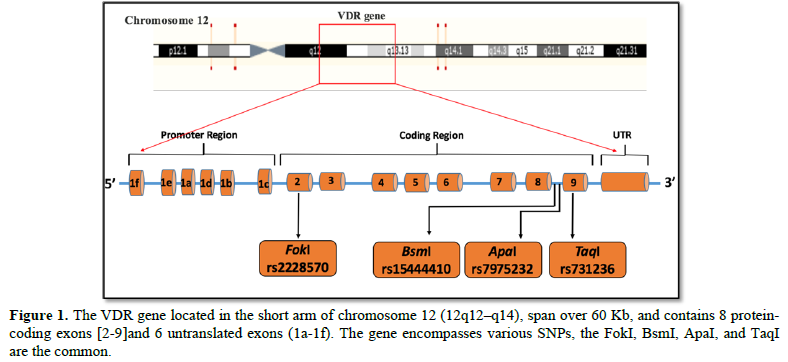
The VDR is the member of the nuclear/steroid receptor superfamily of transcriptional regulations [27], and is found in many tissues in the human body including the islet cells of the pancreas and immune cells, including T lymphocytes and APCs [28]. The human VDR composed of three domains, a C-terminal ligand-binding domain, an N-terminal dual zinc finger DNA-binding domain, and an unstructured region that links the two functional domains [29-31]. This gene spans over 60 kilobases (kb) of genomic DNA and located on the short arm of chromosome 12 (12q12–q14) [29]. The VDR gene contains 14 exons (8 protein-coding exons 2-9, and 6 untranslated exons1a–1f) directed by two distinct promoters [29,30-32]. The 6 untranslated exons were alternatively spliced generating VDR transcripts varied in their 5′ UTRs, and subsequent structural differences in the VDR N-terminal, which may vary between tissues and responsiveness to active vitamin D [29]. The structural VDR gene translated into two functional domains, N-terminal DNA-binding domain (encoded by exon 2 and exon 3) and C-terminal ligand-binding domain (encoded by exon 5-9), linked by unstructured region [31].
The human VDR gene harbors more than 200 SNPs, among them is the big-four-inventory and extensively investigated polymorphisms, FokI F>f alleles (T>C; rs2228570), BsmI B>b (G>A; rs1544410), ApaI A>a (T>G; rs7975232), and TaqI T>t (T>C; rs731236), which were identified with restriction fragment length polymorphism (RFLPs) (Figure 1). The uppercase alleles (F, B, A, or T) represent the absence of the restriction site, while the lowercase alleles (f, b, a, or t) represent the presence of the restriction site.

FokI Polymorphism
FokI polymorphism refers to starting codon polymorphism (ATG) due to thymine (T) to cytosine (C) substitution in the first translational initiation region located in exon 2. The presence of this polymorphism eliminates the original first starting codon (ATG to ACG), and create an alternative start codon six nucleotides downstream of the original. The new start codon leads to translation of mutant protein with an N-terminal three amino acids shorter (C allele or "F" allele; methionine at the fourth position; 424 amino acids) than the wild type protein (T allele or the "f" allele; methionine at the first position; 427 amino acids) [27,33,34]. The polymorphism FokI consequently generates higher transcription rate for the VDR gene and mutant protein 1.5- to 2.5-fold more active than the wild type [35,36]. Pooled and subgroup results from meta-analysis studies showed positive association between FokI polymorphism and the risk of T1DM in Africans [37] and in West Asians [38]. No association has been observed in children [36] or adults in populations from East Asia, Europe, America, Latin America, Australia, and Caucasia [38-44]. Even in the same continent, the results of this polymorphism varied between populations from the same country. For instance, studies in Egyptian children showed significant association of genotypes “ff” with the risk of T1DM [45,46], while others did not report this kind of association [47,48]. High frequency in “Ff” genotype and low of “ff” have been reported in Saudi children with T1DM [49], while other study from the same population doesn’t found this association [50]. Moreover, Iranian studies in adults with T1DM showed discrepancy in results regarding this polymorphism [51,52].
BsmI Polymorphism
BsmI is non-functional polymorphism, located in intron 8, near the 3' end of VDR gene, and correspond to a G to T substitution. This polymorphism has nothing to do with the structure, function, and amount of VDR protein. Nevertheless, and together with ApaI and TagI, the most likely BsmI polymorphic effect on the VDR gene notably arise due to its linkage disequilibrium (LD) with the poly (A) microsatellite repeat in the 3' untranslated region (UTR) [34]. Despite the fact that this latter repeat has inconsistent impact on the VDR mRNA stability and translational activity [53,54], earlier studies have reported strong linkage disequilibrium (LD) between the BsmI alleles and BsmI-ApaI-TaqI haplotypes with the length of the poly A variable number of microsatellites repeat in the 3'UTR [34,55,56].
The proof of association between the non-functional activity of the BsmI polymorphism and the risk of T1DM was controversial among different populations and between continents as well. For instance, pooled results from meta-analysis studies showed positive association of BsmI polymorphism with the risk of T1DM in European, Asian, and Latino populations [39], and with subgroups from East Asia [38,39,43]. In contrary, the polymorphism showed negative association with the risk of T1DM among Americans [37]. Other meta-analysis studies did not find any association in populations from different countries [41,42]. At the allele and genotype levels, "bb" genotype was found to be associated with the risk of T1DM among Asians, Caucasians, Africans, and Latinos, while the “BB” genotype and “B” allele were reported in Africans, Asians, and Latinos [40]. Other meta-analysis study found association between the "Bb" and “bb” genotypes and the risk of T1DM in children from different populations [36].
ApaI Polymorphism
ApaI SNP located in intron 8 at the 3' untranslated region of the gene, and correspond to a T to G transition, which has no functional effect on the VDR protein. Similar to BsmI, ApaI has strong LD with the poly (A) microsatellite repeat in the 3' untranslated region (UTR), particularly it’s allele [27]. Meta-analysis studies among different populations showed no association with this polymorphism and the risk of T1DM [36-44]. However, this SNP showed positive association with the development of T1DM in population from Greece and Pakistan [52,57,58]. On the other hand, SNP low frequency was found in T1DM patients from Saudi Arabia and Egypt [49,59]. This data indicates that this polymorphism alone has little or merely no effect on the risk of T1DM.
TaqI Polymorphism
TagI is a silent SNP (T > C) in axon 9 near the exon-intron boundary (gCTg/ATTg), likely to influence splicing [60]. Like ApaI and BsmI, TagIis strongly linked to a poly(A) microsatellite repeat in the 3’ untranslated region, hence and might affect mRNA stability and the translation of VDR [27,34]. The association of this SNP and the risk of T1DM has be observed in children from different populations, particularly for the normal “TT” and mutant “tt” genotypes [36]. Other meta-analysis studies did not proof an association among adults with T1DM from different continents [41-44]. The lack of association of this SNP with risk of T1DM is attributed to its anonymous effect on the VDR protein, rather than the effect on haplotype and genotype analysis [55,66].
CONCLUSION
The VDR polymorphisms have been extensively studied in relation to different diseases. Of these, the big-four-inventory polymorphisms, FokI, BsmI, ApaI, and TaqI, were broadly investigated between different populations. FokI is only the functional polymorphism, and therefore it has an apparent association with the risk of T1DM, but this association differs in- and between- ethnic groups. The BsmI, ApaI, and TaqI, are nonfunctional polymorphisms, located between intron 8 and exon 9, near the 3' end of VDR gene, and they had strong LD with the poly (A) microsatellite repeat in the 3' untranslated region (UTR) of the VDR gene. Thus far, their plausible association with the risk of T1DM is meant to be hindered, because their anonymous functions still stand against their factual role.
The big-four-inventory polymorphisms and their association with the risk of T1DM have been gathered via consecutive meta-analysis studies. These kinds of studies are conclusive and more reliable as it includes large data sets to evaluate the genotype frequencies in certain diseases, hence reduce the effect of biased sampling that occur in individual study populations. Our review study indicated discrepancies in the association between the aforementioned polymorphisms and the risk of T1DM between continents and ethnics from same populations, when we examined such data from meta-analysis studies. Nevertheless, the hitherto inconclusive data so far is recalling for doing further studies with large sample size and equal geographic distribution of data collections between continents. Performing studies encompassing haplotype’s structure and LD between polymorphisms, together with using advanced techniques other than the RFLP for determining individual polymorphisms, could likely help to emphasize the association of the reviewed SNPs and their linkage with totally different functional alleles that may found in the VDR gene or in genes nearby VDR, and to construct haplotype maps for candidate genes.
CONFLICT OF INTEREST
The author declares no conflicts of interest in this work.
- Richardson SJ, Morgan NG, Foulis AK (2014) Pancreatic pathology in type 1 diabetes mellitus. Endocr Pathol 25: 80-92.
- Atkinson MA, Eisenbarth GS, Michels AW (2014) Type 1 diabetes. Lancet 383: 69-82.
- Todd JA, Walker NM, Cooper JD, Smyth DJ, Downes K, et al. (2007) Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet 39: 857-864.
- IDF Diabetes Atlas Group (2015) Update of mortality attributable to diabetes for the IDF Diabetes Atlas: Estimates for the year 2013. Diabetes Res Clin Pract 109: 461-465.
- Patterson CC, Karuranga S, Salpea P, Saeedi P, Dahlquist G, et al. (2019) Worldwide estimates of incidence, prevalence and mortality of type 1 diabetes in children and adolescents: Results from the International Diabetes Federation Diabetes Atlas, 9th Diabetes Res Clin Pract 157: 107842.
- Verkauskiene R, Danyte E, Dobrovolskiene R, Stankute I, Simoniene D, et al. (2016) The course of diabetes in children, adolescents and young adults: does the autoimmunity status matter? BMC Endocr Disord 16: 61.
- Noble JA, Valdes AM (2011) Genetics of the HLA region in the prediction of type 1 diabetes. Curr Diab Rep 11: 533-542.
- Diaz-Valencia PA, Bougnères P, Valleron AJ (2015) Global epidemiology of type 1 diabetes in young adults and adults: A systematic review. BMC Public Health 15: 255.
- Simmons KM, Michels AW (2015) Type1 diabetes: A predictable disease. World J Diabetes 6: 380-390.
- Ergun-Longmire B, Maclaren NK (2013) Etiology and Pathogenesis of Diabetes Mellitus in Children. In: Feingold KR, Anawalt B, Boyce A, Chrousos G, de Herder WW, et al. (2000) editors Endotext [Internet] South Dartmouth (MA): MDText.com, Inc.
- Vaseghi H, Jadali Z (2016) Th1/Th2 cytokines in Type 1 diabetes: Relation to duration of disease and gender. Indian J Endocrinol Metab 20: 312-316.
- Espinoza-Jiménez A, Peón AN, Terrazas LI (2012) Alternatively activated macrophages in types 1 and 2 diabetes. Mediators Inflamm 2012: 815953.
- Azar ST, Tamim H, Beyhum HN, Habbal MZ, Almawi WY, et al. (1999) Type I (insulin-dependent) diabetes is a Th1- and Th2-mediated autoimmune disease. Clin Diagn Lab Immunol 6: 306-310.
- Rabinovitch A (2003) Immunoregulation by cytokines in autoimmune diabetes. Adv Exp Med Biol 520: 159-193.
- Liblau RS, Singer SM, McDevitt HO (1995) Th1 and Th2 CD4+ T cells in the pathogenesis of organ-specific autoimmune diseases. Immunol Today 16: 34-38.
- Lee YC (2008) Synergistic effect of various regulatory factors in TH1/TH2 balance; immunotherapeutic approaches in asthma. Int J Biomed Sci 4: 8-13.
- Rose K, Penna-Martinez M, Klahold E, Kärger D, Shoghi F, et al. (2013) Influence of the vitamin D plasma level and vitamin D-related genetic polymorphisms on the immune status of patients with type 1 diabetes: a pilot study. Clin Exp Immunol 171: 171-185.
- Dankers W, Colin EM, van Hamburg JP, Lubberts E (2017) Vitamin D in Autoimmunity: Molecular Mechanisms and Therapeutic Potential. Front Immunol 7: 697.
- Kongsbak M, Levring TB, Geisler C, von Essen MR (2013) The vitamin d receptor and T cell function. Front Immunol 4: 148.
- Nykjaer A, Dragun D, Walther D, Vorum H, Jacobsen C, et al. (1999) An endocytic pathway essential for renal uptake and activation of the steroid 25-(OH) vitamin D3. Cell 96: 507-515.
- Jeffery LE, Burke F, Mura M, Zheng Y, Qureshi OS, et al. (2009) 1,25-Dihydroxyvitamin D3 and IL-2 combine to inhibit T cell production of inflammatory cytokines and promote development of regulatory T cells expressing CTLA-4 and FoxP3. J Immunol 183: 5458-5467.
- Tian Y, Wang C, Ye Z, Xiao X, Kijlstra A, et al. (2012) Effect of 1,25-dihydroxyvitamin D3 on Th17 and Th1 response in patients with Behçet's disease. Invest Ophthalmol Vis Sci 53: 6434-6441.
- Overbergh L, Decallonne B, Valckx D, Verstuyf A, Depovere J, et al. (2000) Identification and immune regulation of 25-hydroxyvitamin D-1-alpha-hydroxylase in murine macrophages. Clin Exp Immunol 120: 139-146.
- Korf H, Wenes M, Stijlemans B, Takiishi T, Robert S, et al. (2012) 1,25-Dihydroxyvitamin D3 curtails the inflammatory and T cell stimulatory capacity of macrophages through an IL-10-dependent mechanism. Immunobiology 217: 1292-1300.
- Colin EM, Asmawidjaja PS, van Hamburg JP, Mus AM, van Driel M, Hazes, et al. (2010) 1,25-dihydroxyvitamin D3 modulates Th17 polarization and interleukin-22 expression by memory T cells from patients with early rheumatoid arthritis. Arthritis Rheum 62: 132-142.
- Piantoni S, Andreoli L, Scarsi M, Zanola A, Dall'Ara F, et al. (2015) Phenotype modifications of T-cells and their shift toward a Th2 response in patients with systemic lupus erythematosus supplemented with different monthly regimens of vitamin D. Lupus 24: 490-498.
- Penna-Martinez M, Badenhoop K (2017) Inherited Variation in Vitamin D Genes and Type 1 Diabetes Predisposition. Genes (Basel) 8: 125.
- White JH (2012) Vitamin D metabolism and signaling in the immune system. Rev Endocr Metab Disord 13: 21-29.
- Crofts LA, Hancock MS, Morrison NA (1998) Multiple promoters direct the tissue-specific expression of novel N-terminal variant human vitamin D receptor gene transcripts. Proc Natl Acad Sci U S A 95: 10529-10534.
- Chakhtoura M, Azar ST (2013) The role of vitamin d deficiency in the incidence, progression, and complications of type 1 diabetes mellitus. Int J Endocrinol 2013: 148673.
- Pike JW, Meyer MB (2010) The vitamin D receptor: New paradigms for the regulation of gene expression by 1,25-dihydroxyvitamin D (3). Endocrinol Metab Clin North Am 39: 255-269.
- Mathieu C, Badenhoop K (2005) Vitamin D and type 1 diabetes mellitus: state of the art. Trends Endocrinol Metab 16: 261-266.
- Jurutka PW, Remus LS, Whitfield GK, Thompson PD, Hsieh JC, et al. (2000) The polymorphic N terminus in human vitamin D receptor isoforms influences transcriptional activity by modulating interaction with transcription factor IIB. Mol Endocrinol 14: 401-420.
- Uitterlinden AG, Fang Y, Van Meurs JB, Pols HA, Van Leeuwen JP, et al. (2004) Genetics and biology of vitamin D receptor polymorphisms. Gene 338: 143-156.
- Gross C, Krishnan AV, Malloy PJ, Eccleshall TR, Zhao XY, et al. (1998) The vitamin D receptor gene start codon polymorphism: A functional analysis of FokI variants. J Bone Miner Res 13: 1691-1699.
- Sahin OA, Goksen D, Ozpinar A, Serdar M, Onay H, et al. (2017) Association of vitamin D receptor polymorphisms and type 1 diabetes susceptibility in children: A meta-analysis. Endocr Connect 6: 159-171.
- Zhai N, Bidares R, Makoui MH, Aslani S, Mohammadi P, et al. (2020) Vitamin D receptor gene polymorphisms and the risk of the type 1 diabetes: a meta-regression and updated meta-analysis. BMC Endocr Disord 20: 121.
- Wang G, Zhang Q, Xu N, Xu K, Wang J, et al. (2014) Associations between two polymorphisms (FokI and BsmI) of vitamin D receptor gene and type 1 diabetes mellitus in Asian population: A meta-analysis. PLoS One 9: e89325
- Zhang J, Li W, Liu J, Wu W, Ouyang H, et al. (2012) Polymorphisms in the vitamin D receptor gene and type 1 diabetes mellitus risk: An update by meta-analysis. Mol Cell Endocrinol 355: 135-142.
- Qin WH, Wang HX, Qiu JL, Huang XB, Huang Y, et al. (2014) A meta-analysis of association of vitamin D receptor BsmI gene polymorphism with the risk of type 1 diabetes mellitus. J Recept Signal Transduct Res 34: 372-377.
- Tizaoui K, Kaabachi W, Hamzaoui A, Hamzaoui K (2014) Contribution of VDR polymorphisms to type 1 diabetes susceptibility: Systematic review of case-control studies and meta-analysis. J Steroid Biochem Mol Biol 143: 240-249.
- Guo SW, Magnuson VL, Schiller JJ, Wang X, Wu Y, et al. (2006) Meta-analysis of vitamin D receptor polymorphisms and type 1 diabetes: A HuGE review of genetic association studies. Am J Epidemiol 164: 711-724.
- Wang Q, Xi B, Reilly KH, Liu M, Fu M, et al. (2012) Quantitative assessment of the associations between four polymorphisms (FokI, ApaI, BsmI, TaqI) of vitamin D receptor gene and risk of diabetes mellitus. Mol Biol Rep 39: 9405-9414.
- Ponsonby AL, Pezic A, Ellis J, Morley R, Cameron F, et al. (2008) Variation in associations between allelic variants of the vitamin D receptor gene and onset of type 1 diabetes mellitus by ambient winter ultraviolet radiation levels: A meta-regression analysis. Am J Epidemiol 168: 358-365.
- El-Kafoury AA, Amira MH, Ali ME, Dawoods S (2014) The association of polymorphic sites in some genes with type 1 diabetes mellitus in a sample of Egyptian children. Egypt J Med Hum Genet 15: 265-272.
- Abd-Allah SH, Pasha HF, Hagrass HA, Alghobashy AA (2014) Vitamin D status and vitamin D receptor gene polymorphisms and susceptibility to type 1 diabetes in Egyptian children. Gene 536: 430-434.
- Hamed EO, Abdel-Aal AM, Norel Din AK, Atia MMA (2013) Vitamin D Level and Fok-I Vitamin D Receptor Gene Polymorphism in Egyptian Patients with Type-1 Diabetes. Egypt J Immunol 20: 1-10.
- Ahmed AE, Sakhr HM, Hassan MH, El-Amir MI, Ameen HH, et al. (2019) Vitamin D receptor rs7975232, rs731236 and rs1544410 single nucleotide polymorphisms, and 25-hydroxyvitamin D levels in Egyptian children with type 1 diabetes mellitus: effect of vitamin D co-therapy. Diabetes Metab Syndr Obes 12: 703-716.
- Ali R, Fawzy I, Mohsen I, Settin A (2018) Evaluation of vitamin D receptor gene polymorphisms (Fok-I and Bsm-I) in T1DM Saudi children. J Clin Lab Anal 32: e22397.
- El-Badrawy E, Ibrahim ZS, Aziz AA, Kamel MM, Shehab GM, et al. (2013) Association of Vitamin D Receptors Genes Polymorphism (Apa I, and Taq I) with type 1 diabetes in Saudi Arabia (KSA). Life Science J 10: 1555-1562.
- Bonakdaran S, Abbaszadegan MR, Dadkhah E, Khajeh-Dalouie M (2012) Vitamin D receptor gene polymorphisms in type 1 diabetes mellitus: A new pattern from Khorasan Province, Islamic Republic of Iran. East Mediterr Health J 18: 614-619.
- Mohammadnejad Z, Ghanbari M, Ganjali R, Afshari JT, Heydarpour M, et al. (2012) Association between vitamin D receptor gene polymorphisms and type 1 diabetes mellitus in Iranian population. Mol Biol Rep 39: 831-837.
- Whitfield GK, Remus LS, Jurutka PW, Zitzer H, Oza AK, et al. (2001) Functionally relevant polymorphisms in the human nuclear vitamin D receptor gene. Mol Cell Endocrinol 177: 145-159.
- Durrin LK, Haile RW, Ingles SA, Coetzee GA (1999) Vitamin D receptor 3'-untranslated region polymorphisms: Lack of effect on mRNA stability. Biochim Biophys Acta 1453: 311-320.
- Ingles SA, Haile RW, Henderson BE, Kolonel LN, Nakaichi G, et al. (1997) Strength of linkage disequilibrium between two vitamin D receptor markers in five ethnic groups: Implications for association studies. Cancer Epidemiol Biomarkers Prev 6: 93-98.
- Morrison NA, Qi JC, Tokita A, Kelly PJ, Crofts L, et al. (1994) Prediction of bone density from vitamin D receptor alleles. Nature 367: 284-287.
- Panierakis C, Goulielmos G, Mamoulakis D, Petraki E, Papavasiliou E, et al. (2009) Vitamin D receptor gene polymorphisms and susceptibility to type 1 diabetes in Crete, Greece. Clin Immunol 133: 276-281.
- Mukhtar M, Batool A, Wajid A, Qayyum I (2017) Vitamin D Receptor Gene Polymorphisms Influence T1D Susceptibility among Pakistanis. Int J Genomics 2017: 4171254.
- Kamel MM, Fouad SA, Salaheldin O, El-Razek Ael-R, El-Fatah AI, et al. (2014) Impact of vitamin D receptor gene polymorphisms in pathogenesis of Type-1 diabetes mellitus. Int J Clin Exp Med 7: 5505-5510.
- Hussain T, Naushad SM, Ahmed A, Alamery S, Mohammed AA, et al. (2019) Association of vitamin D receptor TaqI and ApaI genetic polymorphisms with nephrolithiasis and end stage renal disease: A meta-analysis. BMC Med Genet 20:193.
QUICK LINKS
- SUBMIT MANUSCRIPT
- RECOMMEND THE JOURNAL
-
SUBSCRIBE FOR ALERTS
RELATED JOURNALS
- Advances in Nanomedicine and Nanotechnology Research (ISSN: 2688-5476)
- Journal of Veterinary and Marine Sciences (ISSN: 2689-7830)
- Journal of Microbiology and Microbial Infections (ISSN: 2689-7660)
- Proteomics and Bioinformatics (ISSN:2641-7561)
- Journal of Astronomy and Space Research
- Journal of Biochemistry and Molecular Medicine (ISSN:2641-6948)
- Journal of Agriculture and Forest Meteorology Research (ISSN:2642-0449)



