Short Communication
New Data: MHC Genes in Echinodermata
3798
Views & Citations2798
Likes & Shares
MHC genes were recently discovered in 3 classes of Echinodermata out of 5 ones: the Ophuirids, the Crinoïds and more recently: the Asterids. These genes belong to MHC Class I and MHC Class II. Echinids and Holothurids don’t present MHC genes.
INTRODUCTION
MHC genes (Class I, Class II) were discovered, for the first time, in Invertebrates, and particularly in 2 classes of Echinodermata: the ophuirids and the crinoïds out of 5 classes
- They were found more recently in a third class: Asterids
- The aim of this work is to summarize the different obtained results [1,2].
MATERIALS AND METHODS
Three Echinodermata: the Ophuirid (Ophiocomina nigra), the Crinoïd (Antedon bifida) and the Asterid (Asterias rubens) were used.
Their digestive coeca were removed and treated with Trizol (Invitrogen) to obtain RNA.
Transcriptomes from Ophiocomina nigra (Ophuirids) Antedon bifida (Crinoïds) were assembled from RNA-Seq fastq files using Trinity v2.1.1 with default parameters [3]. A BLAST database was created with the assembled transcripts using makeblastdb application from ncbi-blast+ (v2.2.31+). The sequences of transcripts of interest were then blasted against this database using blastn application from ncbi-blast+ with parameter word size 7 [4].
As for Asterias rubens (Asterids), cDNA was normalized using double strand specific nuclease essentially as des cribed by Zhulidov [5]. cDNA was fragmented using DNA Fragmentase (New England Biolabs), according to the manufacturer's instructions. After ligation of adapters for Illumina's GSII sequencing system, the cDNA was sequenced on the Illumina GSII platform sequencing 100 bp from one side of the approximately 200 bp fragments. Sequences were assembled using Velvet Zerbino [6].
Assembled nodes were used for further assembly including Beta vulgaris EST-Data from NCBI in MIRA.
RESULTS
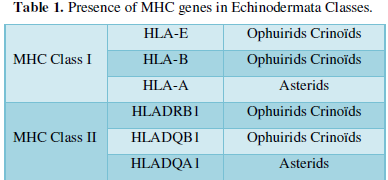
Results are summarized in a table (Table I) which is following. It corresponds to various transcriptomes [1,2] discovered in the 3 classes of Echinodermata with a significant e-value.

CONCLUSION
It appears as clearly as possible that 3 MHC Class I genes are present in Echinodermata: we recall next to HLA-A gene: the HLA-B, the HLA-E genes we described.
On the other hand, 3 MHC Class II genes exist in Echinodermata: HLADRB1, HLADQB1 and HLADQA1 ones.
Further studies are necessary to clarify or to confirm a) the non-existence of MHC Class II genes and Class I in Holothurids which have not axial lymphoid organ and Echinids which have been highly studied in the past, from a genomic point of view.
The Echinids and Holothurids are in fact, the 2 missing classes of Echinodermata which don’t present Adaptative Immunity.
- Leclerc M (2020) Evidence of MHC Class I and Class II Genes in Echinodermata. Proteomics Bioinformatics 2(1): 59-61.
- Leclerc M, Baerlocher L (2021) Mapping on Sea-Star MHC Genes in Invertebrates. Eur J Biol Biotechnol 2(2): 60-62.
- Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, et al. (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnol 29: 644-652.
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3): 403-410.
- Zhulidov PA, Bogdanova EA, Shcheglov AS, Vagner LL, Khaspekov GL, et al. (2004) Simple cDNA normalization using kamchatka crab duplex-specific nuclease. Nucleic Acid Res 3: 32-37.
- Zerbino DR, Birney E (2008) Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18: 821-829.


